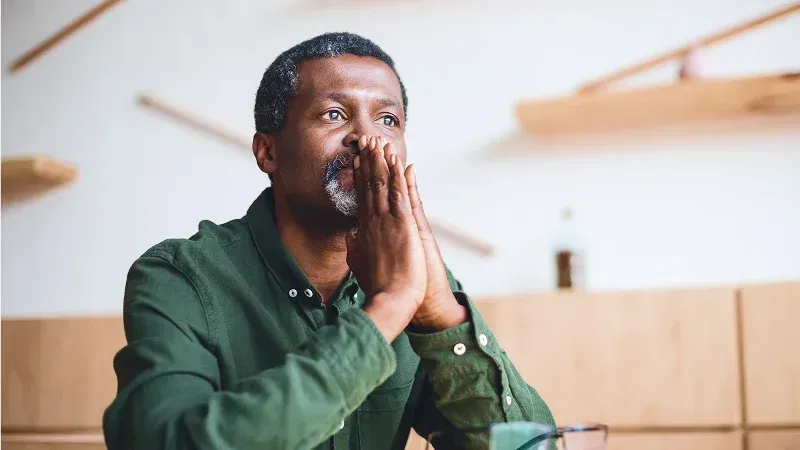

Correcting a misdiagnosis
Identifying risk and finding cancer early is the best way to minimize costs and maximize health and healing. Through Color’s oncologist-led approach we engage your population, driving early detection beyond benchmarks and get them to diagnosis faster, leading to better outcomes and lower costs.
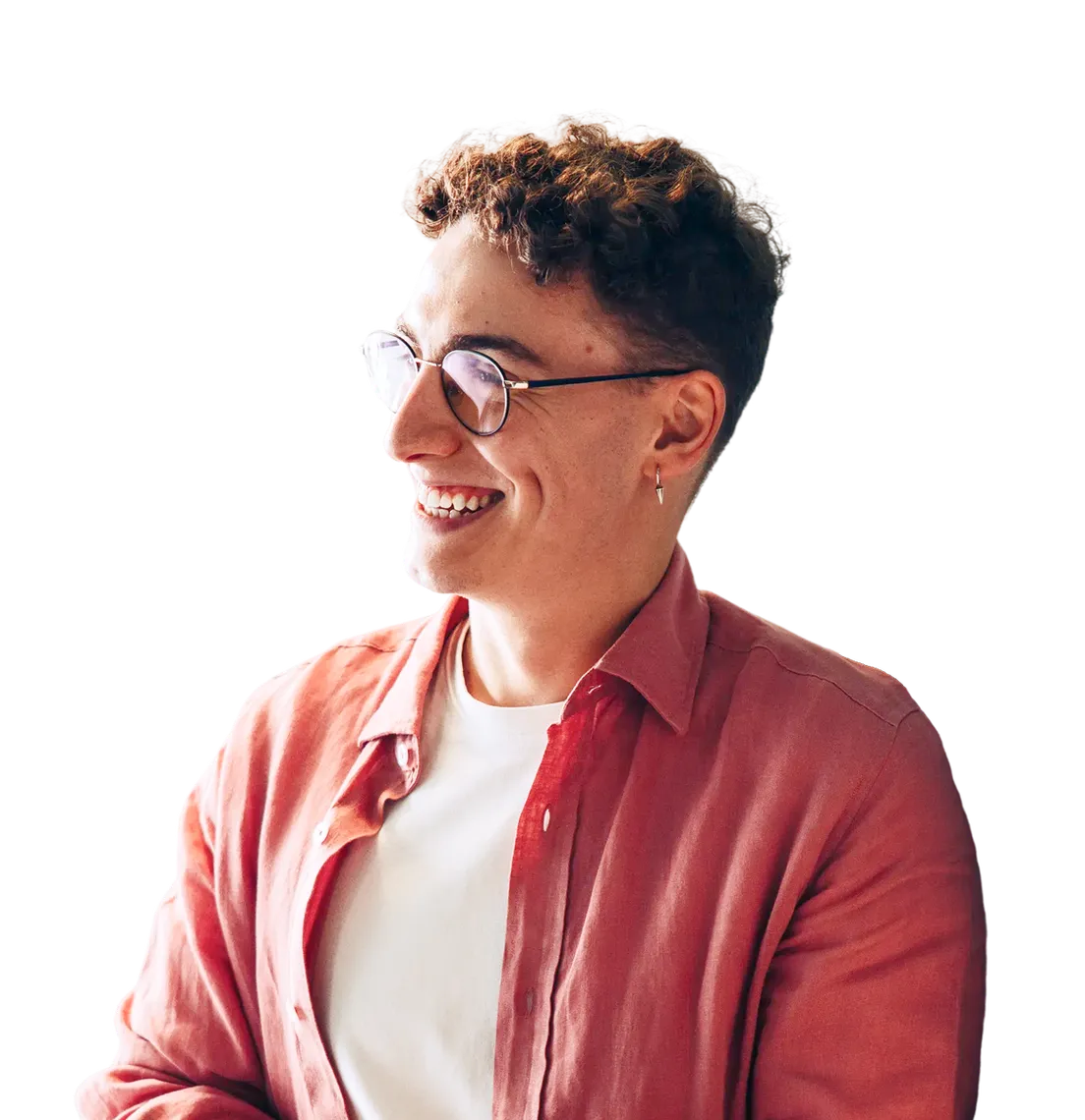

Early detection is the single most powerful lever to improve cancer outcomes and reduce spend. Yet screening is often treated as a one-time task rather than a clinically managed process. Day in and day out follow-ups are missed, and the average cancer diagnoses can take months instead of weeks.
With Color, prevention and early detection become an active, clinician-led strategy that identifies risk sooner, closes care gaps, and gets people to critical answers faster.
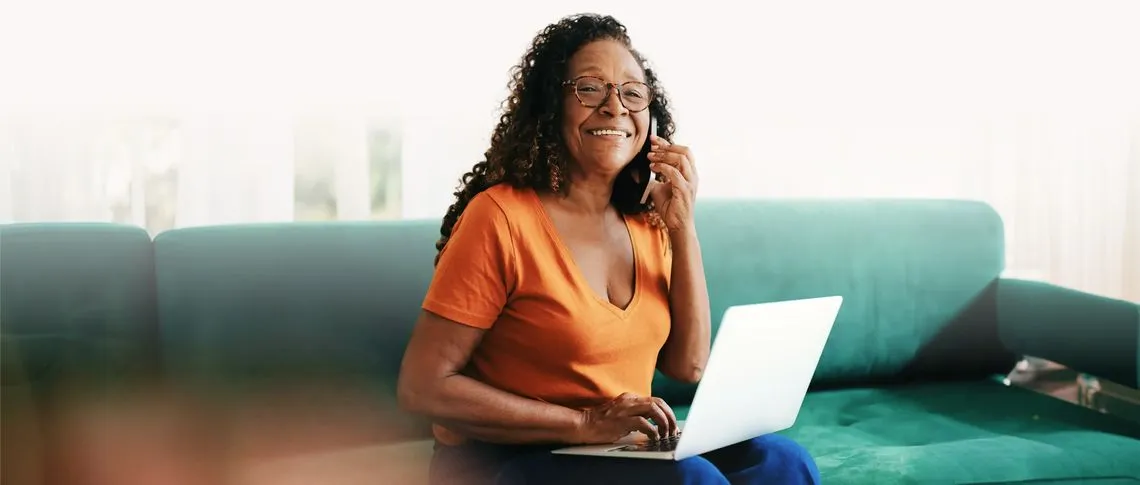

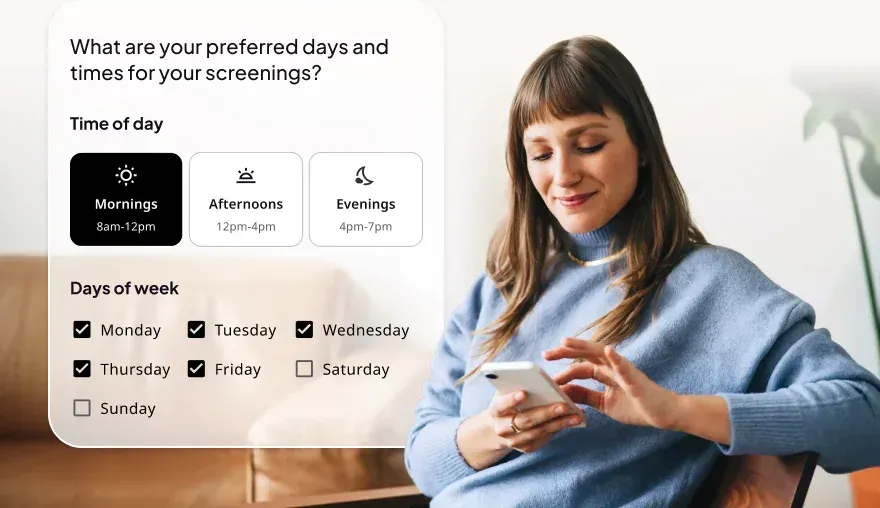
Designed to engage from anywhere, provide access to all members when needed.
Cuts time to diagnosis by 66%, moving patients forward in days, not months.

Physician-licensed care is more effective than simple navigation alone, providing better management and quality of life.

Early Detection & Diagnosis
Planning & Active Treatment →
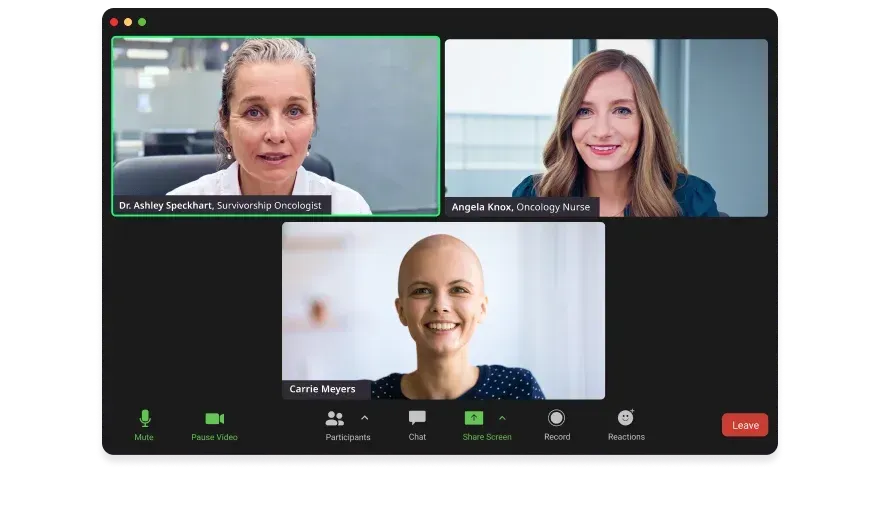
Survivorship Care & Return to Work →
population enrollment in year one
increase in screening adherence
follow-up on abnormal screening results
faster time to diagnosis
reduction in avoidable imaging spend
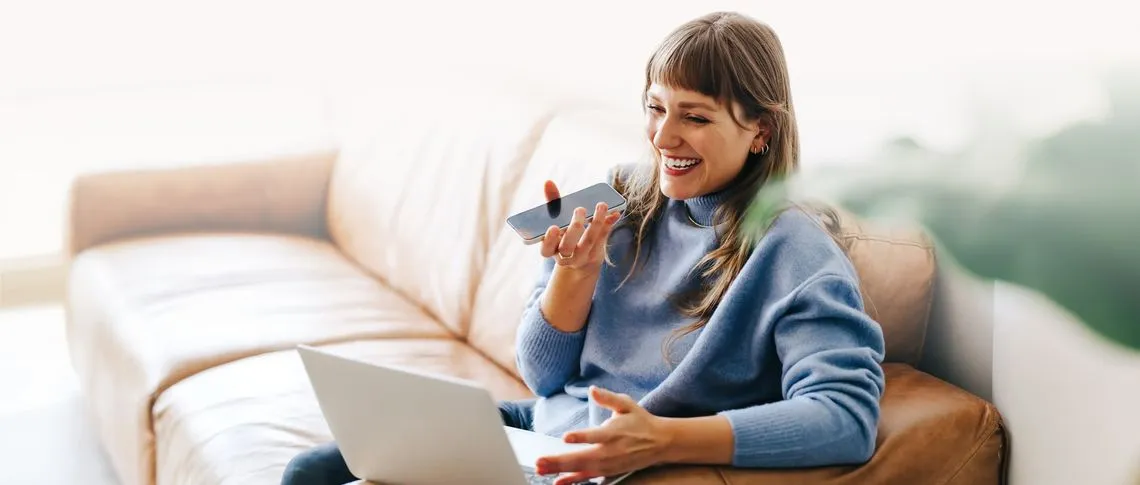

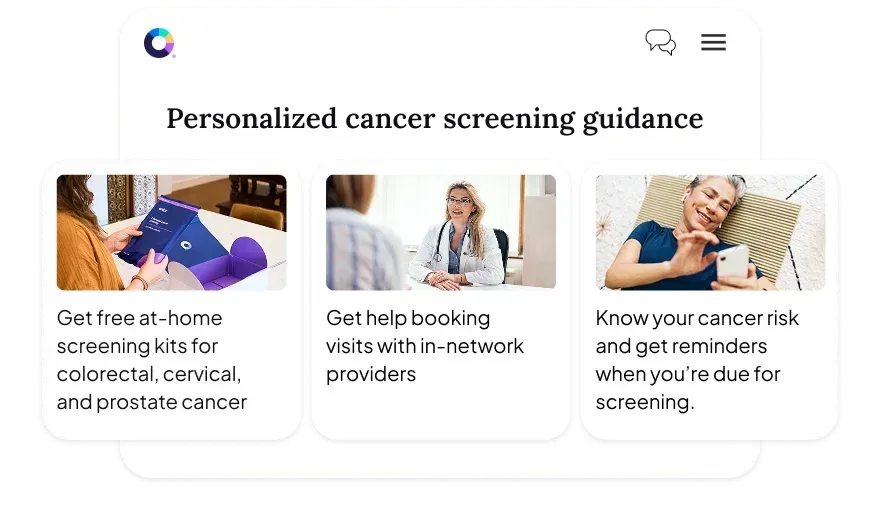
Color starts with a clinical assessment that identifies cancer risk across the population and matches people to the right screening at the right time. This guidelines-based risk assessment goes beyond age or claims data alone to identify higher-risk individuals earlier, reduce unnecessary screening, and shorten time to diagnosis.

Correcting a misdiagnosis
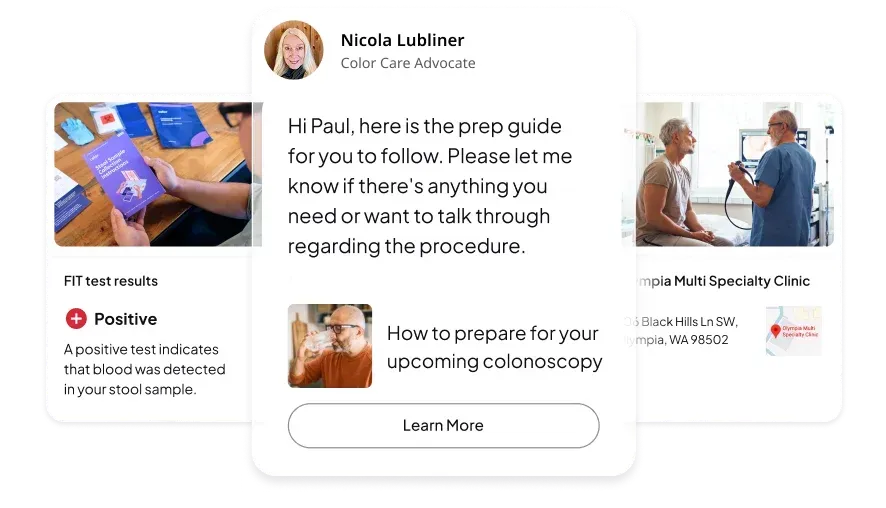
A patient came to Color with a pancreatic cancer diagnosis from their treating provider. After reviewing the case through our tumor board, to assess the diagnosis. Our clinicians identified that the diagnosis was incorrect. It was instead, cholangiocarcinoma, or bile duct cancer, a different disease requiring a completely different treatment plan, prognosis, and clinical trial options.
Because a partnership had been built with the treating provider from day one, Color caught the error before therapy began, avoided unnecessary treatment, and moved the patient quickly into the right care.
Color isn’t like the traditional model. We take action.

Supporting a complex appeal
A woman with dense breast tissue went through the standard prior authorization and appeal process for an MRI, but the clinical documentation did not fully capture the nuances of her risk. She reached out to Color for help understanding her next steps. Our oncologists reviewed her history, completed the necessary risk calculations, and submitted additional clinical details to her health plan as part of the ongoing appeal. With the full context in place, the plan approved the MRI and she advanced quickly into follow-up care.
Color isn’t like the traditional model. We take action.

From postcard to diagnosis in 18 days
A patient hadn't seen a doctor in more than 20 years when she enrolled with Color. During her clinical assessment, our team identified high-risk symptoms that required urgent attention.
Color immediately escalated her to diagnostic imaging, ordered a full workup including biopsy and specialist referral, and confirmed a breast cancer diagnosis in just 18 days, compared to the six-month national average.
Color isn’t like the traditional model. We take action.